Chronic hepatitis C virus affects an around 14 million in Europe and 185 million individuals worldwide. On average, 2-3% of the world’s population is infected with hepatitis C virus (HCV) (Shiffman et al., 2017). Hepatitis C accounted for the greatest number of deaths and the highest mortality rate of 5.0 deaths/100,000 population in 2013 from three types of viral hepatitis (hepatitis A, B, and C). HCV transmission requires that infectious virions contact susceptible cells that allow replication. Most patients (80-85%) who become acutely infected cannot clear the virus and progress to chronic infection (Tyagi, Koirala, 2017). Persistent HCV infection is associated with the development of liver cirrhosis, hepatocellular carcinoma (HCC), liver failure, and death, and a significant portion of liver transplantation in Europe is attributable to disorders related to chronic hepatitis C (CHC) (Petruzziello et al., 2016 ). Chronic HCV infection causes approximately 500,000 deaths each year (Maasoumy et al., 2016 ).
Chronic hepatitis C
Infected population
Mortality
Chronic infection
Understanding the structure of HCV is important because newer therapies target specific viral proteins (Burstow et al., 2017). The Hepatitis C virus is a spherical, enveloped, positive strand RNA virus. It is a member of the family Flaviviridae, yet distinct to be classified as a separate genus, Hepacivirus. The genome is approximately 9.6 kb in length. It encodes a polyprotein that is then posttranslationally modified by proteolytic enzymes into four structural and six nonstructural (NS) proteins. Of particular importance are the NS3/4A, NS5A, and NS5B proteins. The NS3/4A serine protease mediates cleavage of the polyprotein into its respective structural and NS proteins. NS5B is responsible for RNA-dependent RNA polymerase activity, while NS5A is thought to have a number of roles including mediating interferon resistance. Viral proteins work together to drive viral replication and persistence (Burstow et al., 2017 Tyagi, Koirala, 2017).

Hepatitis C virus polyprotein structure (adapted from Burstow et al., 2017)
The most updated classification identified 7 genotypes of Hepatitis C based on their nucleotide variability in HCV sequences recovered from multiple geographic regions (Tyagi, Koirala, 2017).
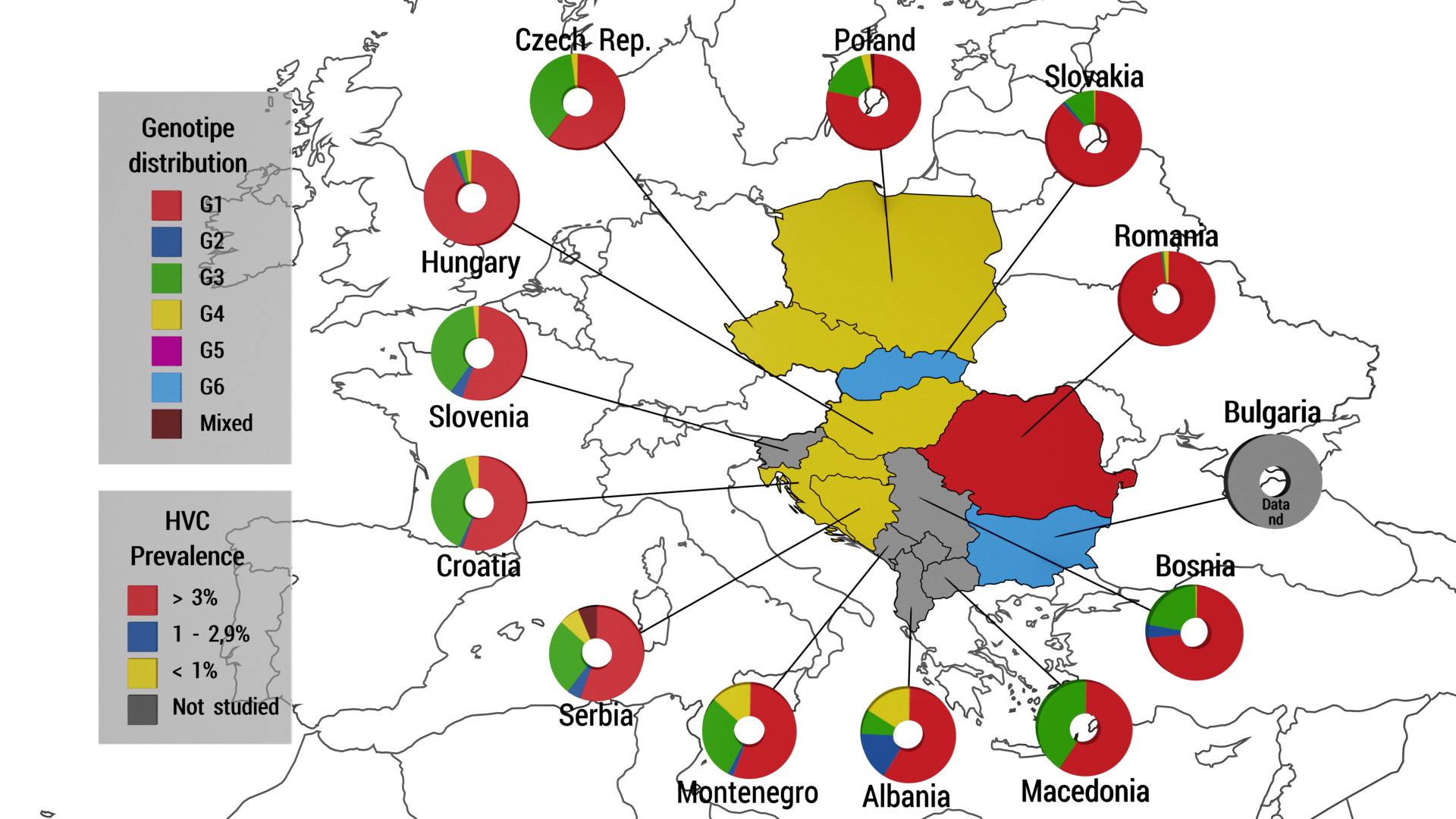
Genotype distribution in Central Europe (adapted from Petruzziello et al., 2016).
The Hepatitis C RNA virus enters the hepatocyte via endocytosis mediated by at least 4 co-receptor molecules. Following internalization in the cytoplasm its positive stranded RNA is uncoated and translated into 10 mature peptides. These are then cleaved by both host proteases and virally encoded proteases known as NS3-4a serine proteases. Mature peptides subsequently go on to reside on the endoplasmic reticulum forming a replication complex, which contains an important enzyme, the NS5B RNA dependent RNA polymerase. This enzyme catalyzes the positive RNA strand into its negative strand intermediate which in turn serves as the template for new positive strand synthesis. These are packaged with core and envelop glycoprotein into mature virions which then exit the cell via exocytosis. Hepatitis C virus has no ability to integrate in the host's genome. The virus can be detected in plasma within days of exposure, often 1 to 4 weeks, viremia peaks in the first 8 to 12 weeks of infection and then plateaus or drops to undetectable levels (viral clearance);in the majority 50-85% it persists. When chronic infection is established, HCV does not appear to be cytopathic; it is the local inflammatory response that triggers fibrogenesis (Tyagi, Koirala, 2017).
HCV infection risk factors include blood transfusion prior to 1992; born to HCV+ mother; current or past intravenous drug use; hemodialysis; high-risk sexual behaviors (particularly unprotected anal intercourse); incarceration; intranasal illicit drug use; unregulated tattoos (Shaffer, Ahuja, 2017). HCV RNA can be detected in blood (including serum and plasma), saliva, tears, seminal fluid, ascitic fluid, and cerebrospinal fluid. Perinatal transmission frequency range from 0% to 4% in larger studies (Tyagi, Koirala,2017). All people with risk factors for hepatitis C should have a serological screening test for antihepatitis C antibodies. These cannot differentiate between past or current HCV infection. A positive screening test should be followed by a test for hepatitis C RNA to confirm the diagnosis. The hepatitis C genotype and viral load should then be determined (Strasser, 2017). Direct testing for HCV RNA is necessary to distinguish between ongoing or prior infection in persons with HCV antibodies (Tyagi, Koirala, 2017).